Mimi's Personal Immune System
Greetings
The age old definition of virus has been "Ultrastructure in size, sufficiently small enough to pass through a bacterial filter". The debate of if virus is living or non living is a never ending one. More recently, new findings in field of virology has brought older definitions into question. For example, very large viruses (large in comparison to other viruses), or now famously known as "Giant viruses" are clearly large enough to be not ultrastructural and wouldn't pass through a bacterial filter. Their genome is nothing like what our databases hold. The debate has stretched to ask, "If Giant Viruses constitute a separate kingdom".
The list of Giant viruses has been growing slowly over recent years. Certain well known examples of Giant viruses include- Pandoravirus, Mamavirus, Mimivirus, Megavirus, Moumouvirus etc. Pandoravirus salinas currently holds the record for maximum size and is calculated to code for 2541 proteins. As already mentioned a huge number of these proteins don't have any homologous sequence in our known database and hence their function cannot be guessed. Interestingly, most of the giant viruses are infective to free living amoeba species.
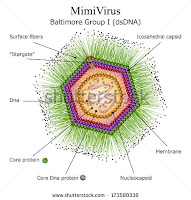 |
| Fig 1: Structure of Mimi Virus. Source |
A paper published yesterday by researchers from France reported that Gigantic mimiviruses has defences similar to the CRISPR system for fighting a virophage called Zamilon. I have a lot of posts dedicated to CRISPR-CAS system in this blog and so will not repeat it here. Interested readers are requested to go through my earlier post.
 |
| Fig 2: The MIMIVIRE defence system. Source |
The research question began with previous reports identifying that not all Mimivirus are susceptible to the Zamilon. They further hypothesised some of them are evolutionarily gained resistance. To test this hypothesis, the team analysed 60 mimivirus genomes and looked for sequences that match those of the Zamilon virophage. This allowed looking for CRISPR-CAS like system. And the team had a clear hit. Mimiviruses that were resistant to Zamilon had a insertion of a repeated Zamilon sequence within an operon, which was named ‘Mimivirus virophage resistance element’ (MIMIVIRE). As Raoult (an author in the study) comments . “They are facing the same kind of challenge that prokaryotes have when they live in communities: they need to fight against viruses and prokaryotes. I even suspect they secrete antibiotic compounds.”
The study investigated the genetic environment of the insertions was compared with CRISPR CAS arrangement. They found important similarities between the 2 systems but also some significant differences. The genetic environment of the insertion had endonuclease (similar to Cas4), helicase etc. As a test of MIMIVIRE gene function systematic gene silencing using siRNA was done. A total of 27 genes were silenced and susceptibility to Zamilon infection was studied. A significant amount of virophage was produced with silencing of R354 (encoding the endonuclease), the R350 gene (encoding helicases and SNF2 domains), and the R349 gene (containing the repeated insert). That makes it pretty much like CRISPR CAS system. The study also did genetic phylogeny analysis using R350 and R354 in an attempt to trace lineage. The genes could be found in other viruses scattered along.
Quoting from Nature news article, “The giant viruses most likely enclose a whole lot of new biology, some of which, including the MIMIVIRE, could find novel application. Maybe in genome editing, maybe in other fields.”
Now I have a lot of questions unanswered- like, does this study also shed some light on possible common ancestor of CRISPR CAS system?
Levasseur, A., Bekliz, M., Chabrière, E., Pontarotti, P., La Scola, B., & Raoult, D. (2016). MIMIVIRE is a defence system in mimivirus that confers resistance to virophage Nature DOI: 10.1038/nature17146
Callaway, E. (2016). CRISPR-like ‘immune’ system discovered in giant virus Nature DOI: 10.1038/nature.2016.19462




Comments
Post a Comment